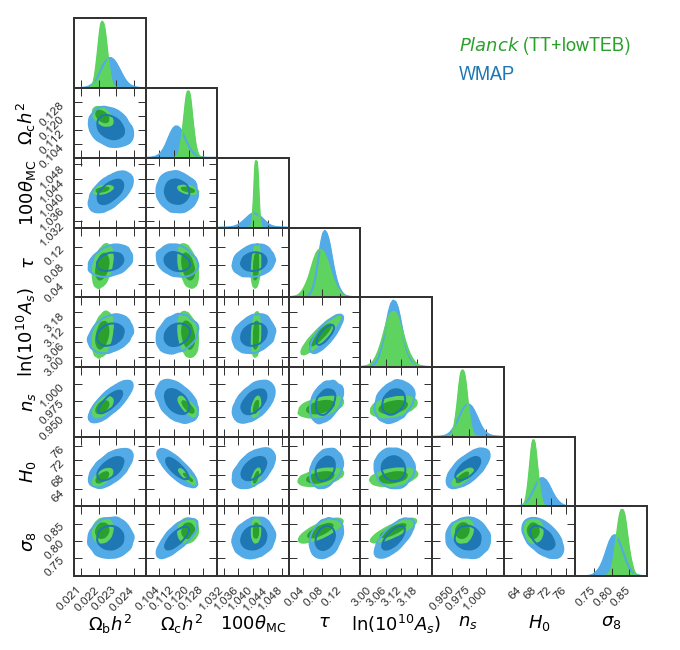
Example 2: Making a GTC/triangle plot with Planck and WMAP data!
This example is built from a jupyter notebook hosted on the pyGTC GitHub repository.
Download the data
The full set of chains from the Planck 2015 release is available at
http://pla.esac.esa.int/pla/#cosmology. You will want to download
COM_CosmoParams_fullGrid_R2.00.tar.gz. Careful, that’s a huge file
to download (3.6 GB)!
Extract everything into a directory, cd into that directory, and run this notebook.
%matplotlib inline
%config InlineBackend.figure_format = 'retina' # For mac users with Retina display
import numpy as np
from matplotlib import pyplot as plt
import pygtc
Read in and format the data
WMAP, Planck = [],[]
for i in range(1,5):
WMAP.append(np.loadtxt('./base/WMAP/base_WMAP_'+str(i)+'.txt'))
Planck.append(np.loadtxt('./base/plikHM_TT_lowTEB/base_plikHM_TT_lowTEB_'+str(i)+'.txt'))
# Copy all four chains into a single array
WMAPall = np.concatenate((WMAP[0],WMAP[1],WMAP[2],WMAP[3]))
Planckall = np.concatenate((Planck[0],Planck[1],Planck[2],Planck[3]))
Select the parameters and make labels
In the chain directories, there are .paramnames files that allow you
to find the parameters you are interested in.
WMAPplot = WMAPall[:,[2,3,4,5,6,7,9,15]]
Planckplot = Planckall[:,[2,3,4,5,6,7,23,29]]
# Labels, pyGTC supports Tex enclosed in $..$
params = ('$\Omega_\mathrm{b}h^2$',
'$\Omega_\mathrm{c}h^2$',
'$100\\theta_\mathrm{MC}$',
'$\\tau$',
'$\ln(10^{10}A_s)$',
'$n_s$','$H_0$',
'$\\sigma_8$')
chainLabels = ('$Planck$ (TT+lowTEB)','WMAP')
Make the GTC!
Produce the plot and save it as Planck-vs-WMAP.pdf.
GTC = pygtc.plotGTC(chains=[Planckplot,WMAPplot],
weights=[Planckall[:,0],
WMAPall[:,0]],
paramNames=params,
chainLabels=chainLabels,
colorsOrder=('greens','blues'),
figureSize='APJ_page',
plotName='Planck-vs-WMAP.pdf')